Modular Network Generation and Visualization Platform with Knowledge Integration Environments
Introduction
Network-based integrative analysis is a powerful technique for deducing biological interpretations from multi-layered omics data such as somatic mutations, copy number variations, and gene expression data. However integrated analysis of multi-omics data is quite complicated and can hardly be done in a fully automated way. Thus, a powerful interactive visual mining tool is much needed in analyzing a huge amount of heterogeneous data such as the TCGA projects.
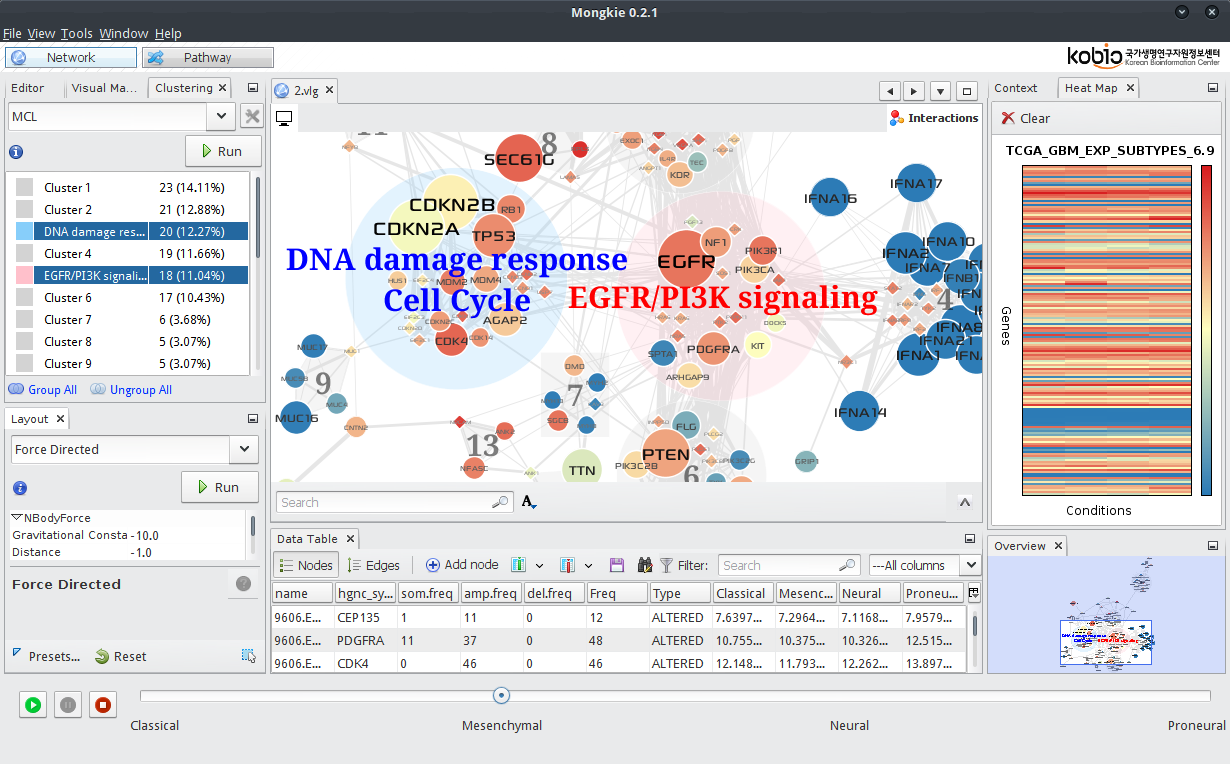
MONGKIE is an application that integrates network visualization with data analysis tools in a single platform. The visualization unit supports sophisticated models for describing biological networks as well as various visualization options (e.g. Data-to-Visual mapping and gene expression overlay) for displaying multi-omics data. In addition, we implemented in-house tools for network analysis including network clustering and over-representation analysis.
Video Demo
Data files used in the demo
- ESC.graphml
- ESC_total_node.csv
- ESC_total_relation.csv
- exp.csv
- GSE15355.csv
- symbol2uniprot.csv (should be imported as a node attribute before the GO enrichment analysis)
Quick Start
See Installation for details.
- Download the latest release of a ZIP distribution for your OS.
- Unzip the downloaded file into any directory.
- Run
mongkie/bin/mongkiein Linux,mongkie\bin\mongkie.exein Windows, andmongkie.app/Contents/MacOS/mongkiein Mac OS X.
What's next? Go to the Tutorial site.
Enjoy!
License
MONGKIE is distributed under the GNU AGPL V3 license with exception of some external libraries that are available under their own licenses.
Visit <http://yjjang.github.io/mongkie> for details.
Copyright (C) 2015 Yeongjun Jang
MONGKIE is free software: you can redistribute it and/or modify
it under the terms of the GNU Affero General Public License as published by
the Free Software Foundation, either version 3 of the License, or
(at your option) any later version.
MONGKIE is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
GNU Affero General Public License for more details.
You should have received a copy of the GNU Affero General Public License
along with this program. If not, see <http://www.gnu.org/licenses/>.